Metabolomics Data Integration and Quality Assessment
Several studies suggest that metabolomics can play a significant role in the investigation of the etiology and progression for many diseases, including cancer. The variety of analytical technologies, laboratory platforms, sample preparation methods, and experimental designs used for metabolic profiling mean that the integration and harmonization of metabolomics data is a difficult challenge. To address these complexities the contractor proposes to develop the Metabolomics Data Integration System (MODIS). MODIS will employ cutting-edge cluster computing technologies to provide bioinformatics methods and database formats for the storage, processing, quality control, and integration of metabolite data across various laboratory platforms and analytical technologies, including LC-MS, GC-MS, and NMR. The development of these methods will position the research community to better leverage existing metabolomics data for the discovery of novel biomarkers and the understanding of the biology of cancer and other diseases.
Publications
Ontology-Based Metabolomics Data Integration with Quality Control, BioAnalysis 2019
Resources
Many of the MODIS components are available in our public GitHub and Docker Hub repositories.
- infotechsoft/htcondor [GitHub | Docker] HTCondor on Docker
- infotechsoft/pegasus-wms [GitHub | Docker] Pegasus Workflow Management System
- infotechsoft/metabolomics [GitHub | Docker] R, XCMS, CAMERA, and Warpgroup tools for MS and metabolomics analysis
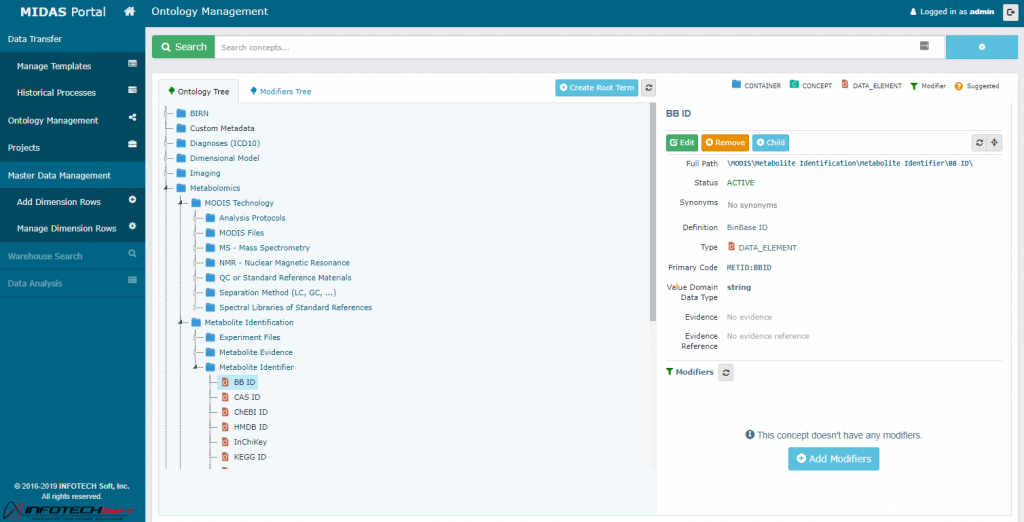
MODIS Platform Demo
For more information about the MODIS platform and Metabolomics Data Integration and Quality Control, please contact us at solutions@infotechsoft.com